HUMAN GLUTATHIONE S-TRANSFERASE A1-1 BINDING WITH NATURALLY OCCURRING LIGANDS: ASSESSMENT BY DOCKING SIMULATIONS
HTML Full TextHUMAN GLUTATHIONE S-TRANSFERASE A1-1 BINDING WITH NATURALLY OCCURRING LIGANDS: ASSESSMENT BY DOCKING SIMULATIONS
Rashi Srivastava * 1, Salman Akthar 2, Rolee Sharma 2 and Sanjay Mishra 1
Department of Biotechnology 1, IFTM University, Moradabad - 244001, Uttar Pradesh, India.
Department of Biotechnology 2, Integral University, Lucknow - 226015, Uttar Pradesh, India.
ABSTRACT: Glutathione S-transferases are phase-II enzymes associated with detoxification and resistance to drugs and xenobiotics. The majority of human tumors and human tumor cell lines express significant amounts of alpha class Glutathione S-Transferase A1-1. Present study aimed to examine hGSTA1-1 interaction with some naturally occurring ligands using docking simulations. Docking simulation using Glutathione S-transferases A1-1 monomer without Glutathione conjugate was receptor against Glutathione, Ellagic acid, Lycopene, α-Tocopherol acetate, Quercetin, Ethacrynic acid, Caffeic acid, Porphyrin, Ferulic acid, Curcumin, Dithiothreitol, Cinnamic acid, Iodoacetamide, α-Tocopherol, Beta-mercaptoethanol showed that the majority ligands tested bound at sites bordering the enzyme subunit-subunit interface. Docking results showed that all the selected ligands docked satisfactorily to the hGSTA1-1 enzyme. Lycopene has a strong binding affinity (Binding energy:-10.68 kcal/mol; docking energy: -15.96 kcal/mol) with hGSTA1-1 amongst selected ligands and predicted as a strong inhibitor against hGSTA1-1. In contrast, investigations using hGSTA1-1 monomer revealed there are additional sites for Ethacrynic acid, Betamercaptoethanol and Glutathione binding rather than H-site as expected from X-ray crystallographic data. In conclusion, the docking simulations suggest that the enzyme subunit interface may be important for hGSTA1-1 interactions with ligands. These findings may provide valuable insights for further research to identify naturally occurring therapeutic agents.
| Keywords: |
Autodock, Glutathione S-Transferase, hGSTA1-1, Molecular docking
INTRODUCTION: Glutathione S-transferases (GSTs) constitute a family of phase II detoxification enzymes that catalyze the conjugation of glutathione with a number of hydrophobic compounds to yield water-soluble derivatives that are excreted in urine or bile 1, 2. The human GST superfamily comprises at least 6 classes of isozymes: alpha, mu, pi, omega, theta, and zeta. 3, 4
Class alpha, mu, and pi GST isoenzymes are overexpressed in rat hepatic preneoplastic nodules and the increased levels of these enzymes are believed to contribute to the multidrug-resistant phenotype observed in these lesions. The majority of human tumors and human tumor cell lines express significant amounts of class alpha, pi, and mu GST.
In recent years, specific GST isoforms have been implicated in the clearance of drugs and environmental toxicants, such as carcinogens and pesticides, as well as in the metabolism of endogenously produced compounds such as lipid peroxidation by-products 4, 5, 6, 7, 8, 9, 10. Many polymorphisms in the genes encoding GSTs have been identified and associated with altered enzyme expression and activity 1, 2. For instance, the GSTM1 gene is deleted in ∼42%–55% in Asians population 11. Therefore, significant inter-individual variation in clearance may exist for any toxicant found to be a substrate of GST.
Furthermore, GSTs have been implicated as one of the causes for drug resistance, especially in cancer, and thus represent potential therapeutic targets 12, 13. For these reasons, a fast and robust method for screening large compound collections against GST isozymes would provide an invaluable tool for identifying compounds liable to interfere with this class of detoxification enzymes.
Human GST A1-1 is a homodimer that belongs to the Alpha class, and each monomer contains 222 amino acids. Several crystal structures of human GST A1-1 have been reported, both of the apo form and various ligand complexes 16, 17. The residues involved in GSH binding at the G site are Tyr9, Arg15, Arg45, Val55, Gln67 and Thr68 from one of the subunits and Asp101 and Arg131 from the other subunit. Deprotonation of GSH contributes significantly to the catalytic activity of GST A1-1 (Ibarra et al., 2003). However, it is not entirely clear why the binding of GSH to the active site lowers the pKa value of the thiol group from 9.2 in solution 15 to 6.7 when bound 18. The conserved residue Tyr9 is positioned within hydrogen-bonding distance of the thiolate group of GSH, and this residue has been shown to be important for the catalytic activity of the enzyme 19. Arg15 is another conserved residue in the Alpha class enzyme that has been shown to stabilize the thiolate ion 20.
Although the complexes with hGST have yet to be studied, there is growing interest in naturally occurring compounds such as tocopherol 21, quercetin 22, ellagic acid and curcumin 23 which inhibit GST. Docking simulations have been successfully applied to model the interaction of GST from filaria 24, 25, 26 and insects 27, 28.
This study aimed to examine hGSTA1-1 interaction with some naturally occurring ligands using docking simulations. To allow for possible receptor-site steric hindrance in the design of this study, we selected single GST subunit for docking simulations. The results are considered regarding binding site motifs and their possible interrelationships with GST activity and structure. This research may provide valuable insights into the effect of naturally occurring compounds on GST and the role of such compounds as chemotherapeutic agents.
MATERIALS AND METHODS: The 3D coordinates of glutathione transferase A1-1 complexed with an ethacrynic acid glutathione conjugate (PDB ID: 1GSE) in Homo sapiens were retrieved from Protein Databank (http://www. rcsb.org/). PROCHECK 32 tool was used to assess the stereochemical properties of the polypeptide structure. The 3D structures for ligands Table 1 were downloaded from the PubChem compound database in SDF format and converted in PDB format with the help of Open Babel 33 tool. All the ligands were subjected to energy minimization and molecular dynamics using the Hyper Chem software 31. Energy calculations were carried out using the AMBER force field. Molecular structure optimization of ligands was carried out using conjugate gradient method Polak-Ribiere until the maximum energy derivative was lower than 0.1kcal/Å mol to obtain a correct geometry.
Docking simulations employed AutoDock v3.0.5 29, 30. Gasteiger charges were added to the ligand, and maximum 6 number of active torsion are given to the lead compound using Auto Dock Tool (http://autodock.scripps.edu/resources/adt).
Kollaman charges and solvation term were added to the protein structure using Auto Dock Tool. The Grid for docking calculation was adjusted to cover entire protein. During the docking procedure, a Lamarckian Genetic Algorithm (LGA) were used for flexible ligand rigid protein docking calculation. Docking parameters were as follows: 30 docking trials, the population size of 150, the maximum number of energy evaluation ranges of 25,0000, the maximum number of generations is 27,000, the mutation rate of 0.02, cross-over rate of 0.8, Other docking parameters were set to the software’s default values.
Modeling operations were performed using a 2GB-64-bit computer to facilitate large 3D rotational grids. The structure for either hGSTA1-1 monomer was loaded to AutoDock keeping default parameters. Shape, electrostatics and potential distributions were used for docking control. Simulations were conducted in the full rotation mode where the ligand is flexible, but the receptor is rigid. Docked poses were analyzed based on the minimum values for the interaction energy. Structures with low values for docking energies were selected, and amino acid residues within 3.5 Å of the ligand were noted as possible binding sites.
TABLE 1: LIST OF LIGANDS
| S. no. | Ligand | CID no. | MWT (g/mol) | Molecular formula | X Log P | HBD | HBA |
| 1 | Glutathione | 124886 | 307.32348 | C10H17N3O6S | -4.5 | 6 | 8 |
| 2 | Ellagic acid | 5281855 | 302.19264 | C14H6O8 | 1.1 | 4 | 8 |
| 3 | Lycopene | 446925 | 536.87264 | C40H56 | 15.6 | 0 | 0 |
| 4 | α-Tocopherol acetate | 2117 | 472.74278 | C31H52O3 | 10.8 | 0 | 3 |
| 5 | Quercetin | 5280343 | 302.2357 | C15H10O7 | 1.5 | 5 | 7 |
| 6 | Ethacrynic acid | 3278 | 303.13798 | C13H12Cl2O4 | 3.8 | 1 | 4 |
| 7 | Caffeic acid | 689043 | 180.15742 | C9H8O4 | 1.2 | 3 | 4 |
| 8 | Porphyrin | 5481276 | 678.7334 | C44H30N4O4 | 5.3 | 6 | 5 |
| 9 | Ferulic acid | 445858 | 194.184 | C10H10O4 | 1.5 | 2 | 4 |
| 10 | Curcumin | 969516 | 368.3799 | C21H20O6 | 3.2 | 2 | 6 |
| 11 | Dithiothreitol | 19001 | 154.251 | C4H10O2S2 | -0.4 | 4 | 4 |
| 12 | Cinnamic acid | 444539 | 148.15862 | C9H8O2 | 2.1 | 1 | 2 |
| 13 | Iodoacetamide | 3727 | 184.96373 | C2H4INO | -0.2 | 1 | 1 |
| 14 | α-Tocopherol | 14985 | 430.7061 | C29H50O2 | 10.7 | 1 | 2 |
| 15 | Betamercaptoethanol | 1567 | 78.13344 | C2H6OS | -0.2 | 2 | 2 |
RESULTS AND DISCUSSION: Glutathione transferases (GSTs) help to protect the cell from potentially toxic alkylating agents that carry electrophilic functional groups, by catalyzing their conjugation with the tripeptide glutathione. The active form of hGSTA1-1 consists of two subunits (Cameron et al., 1995). Each subunit contains two domains: an N-terminal α/β-domain, with βαβαββα topology, and a C-terminal α-helical domain. There are two distinct subsites: a glutathione-binding site (G-site) and a pocket in which the hydrophobic substrates bind (H-site).
According to crystallographic data, 28 interface residues from chain A and 29 interface residues from chain B form the inter-subunit contact (Cameron et al., 1995). The existences of multiple isoforms of GST, together with the relatively non-specific binding of the substrate, meaning that a wide variety of compounds, including certain anti-cancer drugs, can be conjugated to glutathione. Ethacrynic acid (EAA) has been administered to cancer patients in attempts to increase the efficacy of alkylating cytostatic drugs. The rationale is that EAA would serve as GST inhibitors and overcome the resistance caused by GST-dependent inactivation of the cytostatic drug.
Docking simulation using Glutathione S-transferases A1-1 monomer as receptor against natural ligands showed that the majority ligands tested bound at sites bordering the enzyme subunit-subunit interface. Docking results showed that all the selected ligands docked satisfactorily to the hGSTA1-1 enzyme with good docking energies Table 2. Lycopene has a strong binding affinity (Binding energy:-10.68 kcal/mol; docking energy: -15.96 kcal/mol) with hGSTA1-1 amongst selected ligands.
It can be a strong inhibitor against hGSTA1-1. Docked complexes of Glutathione, Ellagic acid, Lycopene, α-Tocopherol acetate, Quercetin, Ethacrynic acid, Caffeic acid, Porphyrin, Ferulic acid, Curcumin, Dithiothreitol, Cinnamic acid, Iodoacetamide, α-Tocopherol, Beta-mercapto-ethanol with hGSTA1-1 were shown in Fig. 2-8.
TABLE 2: ENERGY VALUES OF LIGAND BINDING WITH GLUTATHIONE S-TRANSFERASES MONOMER
| S. no. | Ligand | Binding Energy (Kcal/mol) | Docking Energy
(Kcal/mol) |
Intermol
Energy (Kcal/mol) |
Torsional Energy
(Kcal/mol) |
Internal
Energy (Kcal/mol) |
| 1 | Ellagic acid | -7.87 | -7.85 | -7.87 | 0.0 | 0.02 |
| 2 | Betamercaptoethanol | -3.21 | -3.57 | -3.52 | 0.31 | -0.05 |
| 3 | Glutathione | -1.46 | -5.07 | -4.88 | 3.42 | -0.19 |
| 4 | Lycopene | -10.68 | -15.96 | -15.66 | 4.98 | -0.29 |
| 5 | α-Tocopherol acetate | -3.59 | -8.0 | -7.95 | 4.36 | -0.04 |
| 6 | Quercetin | -8.05 | -7.35 | -8.4 | 0.31 | 1.06 |
| 7 | Ethacrynic acid | -5.78 | -7.81 | -7.65 | 1.87 | -0.17 |
| 8 | Caffeic acid | -5.31 | -5.95 | -5.93 | 0.62 | -0.02 |
| 9 | Porphyrin | -7.16 | -9.84 | -8.41 | 1.25 | -1.43 |
| 10 | Ferulic acid | -5.46 | -6.23 | -6.4 | 0.93 | 0.17 |
| 11 | Curcumin | -7.72 | -7.33 | -10.21 | 2.49 | 2.88 |
| 12 | Dithiothreitol | -3.99 | -5.06 | -4.93 | 0.93 | -0.13 |
| 13 | Cinnamic acid | -5.28 | -5.85 | -5.9 | 0.62 | 0.06 |
| 14 | Iodoacetamide | -4.01 | -4.34 | -4.32 | 0.31 | -0.02 |
| 15 | α-Tocopherol | -4.62 | -9.1 | -8.36 | 3.74 | -0.74 |
The docking studies using a single hGSTA1-1 subunit as template showed a variety of binding sites Table 3.
TABLE 3: LIGAND BINDING SITE IDENTIFIED USING HUMAN GLUTATHIONE S-TRANSFERASE A1-1 MONOMER AS DOCKING TEMPLATE
| S. no. | Ligand | Amino acid residues |
| 1 | Ellagic acid | GLU104, HIS159, LEU163, GLU162, GLY103, LEU107, LYS15, TYR166 |
| 2 | Beta-mercaptoethanol | GLY201, SER202, ARG204, PRO200, GLN199, LEU198 |
| 3 | Glutathione | GLY14, ARG13, LEU107, SER18, LEU72, ALA100, GLY103, ILE106, GLU162, TYR166, ARG155, HIS159, LEU163, ILE96, ILE99 |
| 4 | Quercetin | GLU17, TYR166, GLY14, MET16, SER18, GLU162, ARG15, HIS159, LEU72, ARG155, GLU104, ALA100, ILE96, ARG69, GLU97 |
| 5 | Curcumin | ALA100, ILE99, HIS159, GLY103, ILE106, TYR166, GLU17, ARG13, GLY14, GLU169, LEU170 |
| 6 | Ferulic acid | GLY14, MET208, ARG13, LEU107, GLY103, ILE106 |
| 7 | Caffeic acid | ARG15, SER18, HIS159, LEU107, GLU162, LEU163, GLU104 |
| 8 | Cinnamic acid | PRO207, PRO206, ARG13, ARG204, SER202, GLY201, GLN199, LYS205 |
| 9 | Lycopene | GLN54, VAL55, GLN67, THR68, ARG15, ARG69, LEU107, GLY103,
ARG13, PRO207, GLU169, LEU170, SER172 |
| 10 | α-Tocopherol acetate | ARG45, GLN54, GLN67, ARG15, TYR9, PHE220, VAL55 |
| 11 | Ethacrynic acid | PHE30, VAL28, GLY27, ALA24, SER202, GLY201, LYS196, PRO200 |
| 12 | Iodoacetamide | SER202, ARG204, PHE197, GLN199, LEU198 |
| 13 | α-Tocopherol | ASP209, MET208, PRO110, ARG13, PRO207, LEU109, TYR166, ILE106, GLU169, LEU170 |
| 14 | Dithiothreitol | GLY14, GLU17, ARG13, SER18, LEU107, GLU162, TYR166 |
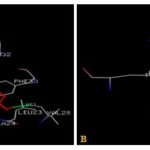
| 15 | Porphyrin | PHE30, VAL28, LEU23, ALA24, LYS196, PRO203, GLY201, PRO200, GLN199, GLU32 |
Residues within a 3.5Å radius of the binding site for the Ethacrynic acid with subunit A of hGSTA1-1 included PHE30, VAL28, GLY27, ALA24, SER202, GLY201, LYS196, PRO200 Fig. 2a rather than H-site (MET208, ARG13, LYS15, GLY14, LEU107, TYR9, PHE222) as expected from X-ray data shown in Fig. 1a.
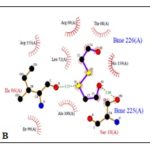
FIG. 1(A-B-C): (A): SCHEMATIC ILLUSTRATING THE RESIDUES CONTACTING ETHACRYNIC ACID (EAA). (B): RESIDUES CONTACTING BETA-MERCAPTOETHANOL (BME). (C): RESIDUES CONTACTING GLUTATHIONE (GSH). THESE FIGURES WERE TAKEN FROM PDBSUM ENTRY OF 1GSE
Binding site residues within a 3.5Å radius of Beta-mercaptoethanol with subunit A of hGSTA1-1 included GLY201, SER202, ARG204, PRO200, GLN199, LEU198 Fig. 2b rather than H-site (ARG155, ARG69, THR68, LEU72, HIS159, ILE96, ALA100, SER18) as expected from X-ray data shown in Fig. 1b.
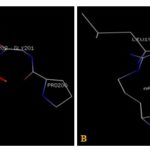
FIG. 2(A-B): (A): THE ETHACRYNIC ACID BINDING SITE RESIDUES. (B): THE BETA-MERCAPTOETHANOL BINDING SITE RESIDUES. THESE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 (PDB ID: 1GSE) MONOMER AS DOCKING TEMPLATE. LIGANDS ETHACRYNIC ACID AND BETA-MERCAPTOETHANOL SHOW IN STICK & BALL REPRESENTATION AND IS COLORED WITH RED USING PYTHON MOLECULAR VIEWER. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY THE GREEN LINE
Residues within a 3.5Å radius of the binding site for the Glutathione with subunit A of hGSTA1-1 included GLY14, ARG13, LEU107, SER18, LEU72, ALA100, GLY103, ILE106, GLU162, TYR166, ARG155, HIS159, LEU163, ILE96, ILE99 Fig. 3a rather than H-site (ASP101, GLN67, THR68, PRO56, TYR9, GLN54, VAL55, ARG131, ARG45, PHE220) as expected from X-ray data shown in Fig. 1c.
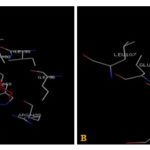
FIG. 3(A-B): (A): THE GLUTATHIONE BINDING SITE RESIDUES.(B): THE ELLAGIC ACID BINDING SITE RESIDUES. THESE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 (PDB ID: 1GSE) MONOMER AS DOCKING TEMPLATE. LIGANDS GLUTATHIONE AND ELLAGIC ACID ARE SHOWN IN THE STICKS AND IS COLORED WITH RED. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY A DOTTED YELLOW LINE AND BY THE GREEN LINE
A possible explanation for this result is that using the enzyme monomer without Glutathione conjugate as docking template leads to unreliable energy minima. As support of this view, in the structure of the complex with EAA, Beta-mercaptoethanol and Glutathione substrate are revealed as binding in a non-productive mode, suggesting that the substrate will only form an active complex when glutathione is already bound. This is biologically sensible as the intracellular levels of glutathione are much higher than the dissociation constant of its complex with GST Al-1. Glutathione is, therefore, probably involved in the molecular recognition of the electrophilic substrate and should be considered when designing drugs to inhibit GST.
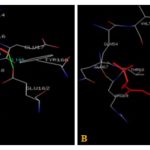
FIG. 4(A-B): (A): THE QUERCETIN BINDING SITE RESIDUES. (B): LYCOPENE BINDING SITE RESIDUES. THESE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 (PDB ID: 1GSE) MONOMER AS DOCKING TEMPLATE. LIGAND QUERCETIN AND LYCOPENE ARE SHOWN IN STICKS REPRESENTATION AND IS COLORED WITH RED USING PYTHON MOLECULAR VIEWER. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY THE GREEN LINE
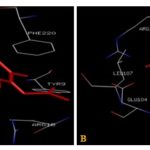
FIG. 5(A-B): (A): THE Α-TOCOPHEROL ACETATE BINDING SITE. (B): THE CAFFEIC ACID BINDING SITE RESIDUES. THESE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 (PDB ID: 1GSE) MONOMER AS DOCKING TEMPLATE. LIGAND Α-TOCOPHEROL ACETATE AND CAFFEIC ACID ARE SHOWN IN STICKS AND STICKS & BALL REPRESENTATION AND IS COLORED WITH RED USING PYTHON MOLECULAR VIEWER. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY THE GREEN LINE
FIG. 6(A-B): (A): THE PORPHYRIN BINDING SITE RESIDUES. (B): THE DITHIOTHREITOL BINDING SITE RESIDUES. THESE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 (PDB ID: 1GSE) MONOMER AS DOCKING TEMPLATE. LIGAND PORPHYRIN AND DITHIOTHREITOL ARE SHOWN IN STICKS REPRESENTATION AND IS COLORED WITH RED USING PYTHON MOLECULAR VIEWER. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY THE GREEN LINE
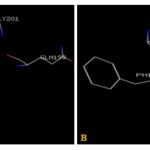
FIG. 7(A-B): (A): THE CINNAMIC ACID BINDING SITE RESIDUES. (B): THE IODOACETAMIDE BINDING SITE RESIDUES. THESE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 (PDB ID: 1GSE) MONOMER AS DOCKING TEMPLATE. LIGAND CINNAMIC ACID AND IODOACETAMIDE ARE SHOWN IN STICKS AND STICKS & BALL REPRESENTATION AND IS COLORED WITH RED USING PYTHON MOLECULAR VIEWER. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY THE GREEN LINE
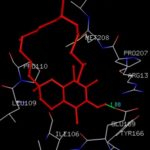
FIG. 8: THE Α-TOCOPHEROL BINDING SITE RESIDUES ARE IDENTIFIED USING GLUTATHIONE S-TRANSFERASE A1-1 MONOMER AS DOCKING TEMPLATE. LIGAND Α-TOCOPHEROL IS SHOWN IN STICKS REPRESENTATION AND IS COLORED WITH RED USING PYTHON MOLECULAR VIEWER. AMINO ACID RESIDUES ARE A REPRESENTATION AS LINES. A HYDROGEN BOND IS REPRESENTED BY THE GREEN LINE
CONCLUSION: Single enzyme subunit was found to provide a wider range of binding sites for naturally occurring ligands. Some inherent limitations of docking were considered perhaps explaining the low level of agreement between docking interaction energies and observed experimental values for enzyme-ligand binding. Finally, the evidence is also presented for differential ligand binding to the GST monomer which, if confirmed by direct measurements, could open the intriguing possibility that the single hGSTA1-1 subunit might be formed under some physiological conditions. More research is needed to determine whether naturally occurring ligands can disturb the structure and stability of hGSTA1-1.
ACKNOWLEDGEMENT: Nil
CONFLICT OF INTEREST: Nil
REFERENCES:
- Eaton DL and Bammler TK: Concise review of the glutathioneS-transferases and their significance to toxicology. Toxicol Sci 1999; 49: 156-164.
- Hayes JD, Flanagan JU and Jowsey IR: Glutathione transferases.Annu Rev Pharmacol Toxicol 2005; 45: 51-88.
- Townsend D and Tew K: Cancer drugs, genetic variation, and the glutathione-S-transferase gene family.Am J Pharmacogenomics 2003; 3: 157-172.
- Mathew N: GlutathioneS-transferase (GST) inhibitors. Expert Opin 2006; 16: 431-444.
- Abel EL, Bammler TK and Eaton DL: Biotransformation of methyl parathion by glutathione S-transferases.Toxicol Sci 2004; 79: 224-232.
- Hamilton DS, Zhang X, Ding Z, Hubatsch I, Mannervik B and Houk KN: Mechanism of the glutathione transferase-catalyzed conversion of antitumor 2-crotonyloxymethyl-2-cycloalkenones to GSH adducts.J Am Chem Soc 2003; 125: 15049-15058.
- Hu X, O’Donnell R, Srivastava SK, Xia H, Zimniak P and Nanduri B: Active site architecture of polymorphic forms of human glutathioneS-transferase P1-1 accounts for their enantioselectivity and disparate activity in the glutathione conjugation of 7beta,8alpha-dihydroxy-9alpha,10alpha-oxy-7,8,9,10-etrahydrobenzo(a) pyrene. Biochem Biophys Res Commun 1997; 235: 424-428.
- Hubatsch I, Ridderström M and Mannervik B: Human glutathione transferase A4-4: an alpha class enzyme with high catalytic efficiency in the conjugation of 4-hydroxynonenal and other genotoxic products of lipid peroxidation.Biochem J 1998; 330((Pt 1)): 175-179.
- Zimniak P, Singhal SS, Srivastava SK, Awasthi S, Sharma R and Hayden JB: Estimation of genomic complexity, heterologous expression, and enzymatic characterization of mouse glutathioneS-transferase mGSTA4-4 (GST 5.7). J Biol Chem 1994; 269: 992-1000.
- Lien S, Larsson AK and Mannervik B: The polymorphic human glutathione transferase T1-1, the most efficient glutathione transferase in the denitrosation and inactivation of the anticancer drug 1, 3-bis(2-chloroethyl)-1-nitrosourea.Biochem Pharmacol 2002; 63: 191-197.
- Gao Y, Cao Y, Gao F and Jian L: An updating meta-analysis of the GSTM1, GSTT1, and GSTP1 polymorphisms and prostate cancer: a HuGE review. Prostate 2009; 69: 662-688.
- Riggs PK, Angel JM, Abel EL and DiGiovanni J: Differential gene expression in the epidermis of mice sensitive and resistant to phorbol ester skin tumor promotion.Mol Carcinog 2005; 44: 122-136.
- TownsendD and Tew K: Cancer drugs, genetic variation, and the glutathione-S-transferase gene family. Am J Pharmacogenomics 2003; 3: 157-172.
- Ibarra CA, Chowdhury P, Petrich JW and Atkins WM: The anomalous pKa of Tyr-9 in glutathione S-transferase A1-1 catalyzes product release. J Biol Chem 2003; 278(21): 19257-19265.
- Jung G, Breitmaier E and Voelter W: Dissoziations gleichgewichte von Glutathion. J. Biochem 1972; 24(3): 438-445.
- Sinning I, Kleywegt GJ, Cowan SW, Reinemer P, Dirr HW, Huber R, Gilliland GL, Armstrong RN, Ji X and Board PG: Structure determination and refinement of human class alpha glutathione transferase A1-1, and a comparison with the Mu and Pi class enzymes. J Mol Biol 1993; 232: 192-212.
- Cameron AD, Sinning I, L'Hermite G, Olin B, Board PG, Mannervik B and Jones TA: Structural analysis of human alpha-class glutathione transferase A1-1 in the apo-form and complexes with ethacrynic acid and its glutathione conjugate. Structure 1995; 3: 717-727
- Gustafsson A and Mannervik B: Benzoic acid derivatives induce recovery of catalytic activity in the partially inactive Met208Lys mutant of human glutathione transferase A1-1. J Mol Biol 1999; 288: 787-800.
- Stenberg G, Board PG and Mannervik B: Mutation of an evolutionarily conserved tyrosine residue in the active site of a human class Alpha glutathione transferase. FEBS Lett 1991; 293(1-2): 153-155.
- Bjo¨ rnestedt R, Stenberg G, Widersten M, Board PG, Sinning I, Jones TA and Mannervik B: Functional significance of arginine 15 in the active site of human class alpha glutathione transferase A1-1. J Mol Biol 1995; 247(4): 765-773.
- Van Haaten RIM, Evelo CTA, Penders J, Eijnwa-chter MP, Haenen GR and Bast A: Inhibition of human glutathione S-transferase P1-1 by tocopherols and α-tocopherol derivatives. Biochimica Biophysical Ac- ta-Protein Structure and Molecular Enzymology 2001; 1548: 23-28.
- van Zanden JJ, Ben Hamman O, van Lersel MLPS, Boeren S, Cnubben NHP, Bellod ML, Vervoort J, van Bladerena PJ and Rietjensa IMCM: Inhibition of human glutathione S-transferase P1-1 by the flavonoid quercetin. Chemico-Biological Interactions 2003; 145: 39-148.
- Hayeshi R, Mutingwende I, Mavengere W, Masiyanise V and Mukanganyama S: The inhibition of human glutathione S-transferases activity by plant poly-phenolic compounds ellagic acid and curcumin. Food and Chemical Toxicology 2007; 45: 286-295.
- Nathan ST, Mathew N, Kalyanasundaram M and Balaraman K: Structure of glutathione S-trans- ferase of the filarial parasite Wuchereria bancrofti: A target for drug development against the adult Journal of Molecular Modelling 2005; 11: 194-199.
- Yadav M, Singh A, Rathaur S and Liebau E: Structural modeling and simulation studies of Brugia malayi glutathione-S-transferase with compounds exhibiting anti-filarial activity: Implications in drug targeting and designing. Journal of Molecular Graphics and Modeling, 2010; 28:435-445.
- Srinivasan L, Mathew N, Muthuswamy K: In-vitro anti-filarial activity of glutathione S-transferase inhibitors. Parasitology Research 2010; 105: 1179-1182.
- Kapoli P, Axarli IA, Platis D, Fragoulaki M, Paine M, Hemingway J, Vontas J and Labrou NE: Engineering sensitive glutathione transferase for the detection of xenobiotics. Biosensors and Bioelectronics 2008; 24: 498-503.
- Setzer WN: The molecular mechanism for DDT detoxification in Anopheles gambiae: A molecular dock-ing Journal of Biophysical Chemistry 2010; 2: 135-136.
- Goodsell DS and Olson AJ: Automated docking of substrates to proteins by simulated annealing. Proteins Structure, Function Bioinformatics 1990; 8: 195-202.
- Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK and Olson AJ: Automated docking using a lamarckian genetic algorithm and empirical binding free energy function. J. Computational Chem 1998; 19: 1639-1662.
- Hyper Chem (TM) Release 7.5, Hypercube, Inc., 1115 NW 4th Street, Gainesville, Florida 32601, USA.
- Laskowski RA, MacArthur MW, Moss DS and Thornton JM: PROCHECK: A program to check the stereochemical quality of protein structures. J App Cry 1993; 26: 283-291.
- O'Boyle NM, Banck M, James CA, Morley C, Vandermeersch T and Hutchison GR: Open Babel: An open chemical toolbox. Journal of Cheminformatics 2011;3: 33.
How to cite this article:
Srivastava R, Akthar S, Sharma R and Mishra S: Human glutathione s-transferase A1-1 binding with naturally occurring ligands: assessment by docking simulations. Int J Pharmacognosy 2014; 1(7): 457-64. doi: 10.13040/IJPSR.0975-8232.1(7).457-64.
This Journal licensed under a Creative Commons Attribution-Non-commercial-Share Alike 3.0 Unported License.
Article Information
9
457-464
1224
2110
English
IJP
R. Srivastava *, S. Akthar, R. Sharma and S. Mishra
Department of Biotechnology, IFTM University, Moradabad, Uttar Pradesh, India.
talk2rashii@gmail.com
14 March 2014
08 May 2014
28 June 2014
http://dx.doi.org/10.13040/IJPSR.0975-8232.IJP.1(7).457-64
01 July 2014