ASSOCIATION STUDY OF YIELD AND ITS COMPONENTS IN PEA (PISUM SATIVUM L.)
HTML Full TextASSOCIATION STUDY OF YIELD AND ITS COMPONENTS IN PEA (PISUM SATIVUM L.)
Natalia Georgieva * 1, Ivelina Nikolova 1 and Valentin Kosev 2
Department of Technology and Ecology of Forage Crops 1, Department of Breeding of Forage Crops 2, Institute of Forage Crops, Pleven, Bulgaria.
ABSTRACT: Field pea is the third most important grain legume globally after beans and chickpea. The present study was conducted during the period 2012-2014 at the Institute of Forage crops, Bulgaria, with five pea genotypes. The conducted analyses revealed that the genetic correlations for most of the traits in field pea have higher coefficients than the phenotypic correlations. The strongest positive phenotypic correlations were found between plant height with pods per plant (r=0.780); pods per plant with seed per plant (r=0.863) and seed weight per plant (r=0.796); seed per plant with seed weight per plant (r=0.733). The highest genetic correlations were established between plant height with seed per plant(r = 0.612) and pods per plant (r=0.798); pods per plant with seed per plant (r=0.866) and seed weight per plant (r=0.796); seed per plant with seed weight per plant (r=0.722). The strongest and direct positive effects over grain yield had pod stem (2991.134), seeds per plant (448.314) and pods per plant (99.261). Based on the trait association and the path coefficients for seed yield ant its components, it could be concluded that pea breeders should pay attention to the traits as pod stem, seeds per plant, pods per plant and 1000 seed weight at selecting high-yielding genotypes.
| Keywords: |
Pisum sativum, Breeding, Correlations, Path analysis
INTRODUCTION: Field pea is high yielding legume crop that is grown worldwide, believed to be the native of the Mediterranean region of Southern Europe and Western Asia. It is the third most important grain legume globally after beans and chickpea 12. Field pea genotypes are distinguished with a great variety of morphological, phenological and economic traits which are used as indicators in characterization and differentiation of accessions, hybrids, and lines in this crop.
Between some traits of the genotypic characteristic is established that there are correlations that allow in the expression of one of the parameters to prognosticate the manifestation of the trait determined by the same genetic mechanism.
Before initiating any effective selection program it is necessary to know the importance and association of various components with yield and among each other as unfavorable association between the desired attributes under selection may result genetic slippage and limit the genetic advance and the yield being the end product of many correlated characters, the selection of this character would be more effective when it is based on component characters which are highly heritable and positively correlated. When number of variables is considered in correlation, the association becomes more and more complex 3. The efficiency of selection based on estimates of character association can further be enhanced by the as curtaining direct and indirect effect of component traits towards an expression of seed yield 4.
The scientific researches in the field of pea selection marked significant progress in creating new varieties with higher nutritional value, improved environmental plasticity and stability 5. Despite the great success in this area need and opportunities for further improvement and creation of new pea varieties are still a major task due to constantly changing growing conditions and the lack of suitable varieties for these conditions 6.
In the genetic-selection researches for analyzing the causal link in systems of correlating magnitudes are used different criteria and analyses – correlation coefficients, multiple regression and Pat-analysis 7. Coefficients of Path-analysis can be used for assessment of interdependencies between the yield and yield components, and they are more effective than the other methods 8. This method is gaining ever more ground in biological researches due to the possibility of disclosure of direct and indirect influence of the traits.
Information of the variability and inter-dependencies of quantitative traits in plant population is essential to increase the efficiency of breeding 9. Present study aimed to determine the relationships among seed yield components of spring pea cultivars using correlation and path coefficient analysis.
MATERIALS AND METHODS: The experi-mental work was carried out from 2012 to 2014 at the Institute of Forage Crops, Pleven (43.41ºN, 24.61ºE), situated in the central part of the Danube hilly plain. Five spring pea (Pisum sativum L.) cultivars were included in the trial namely Glyans, Svit, Kamerton, and Modus from the Ukraine and Pleven 4 from Bulgaria. The long plot design was used where plot size was 4 m2. The sowing was done by hand, at a depth of 4 cm and with a rate of 120 seeds per m2. The plants were cultivated in conditions of organic farming (without the use of any fertilizers and pesticides). For seed yield (kg da-1) data for each cultivar was recorded on the basis of seed yield per plot at the end of growing season. Seed yield in each cultivar was measured at a standard seed humidity of 14%.
Ten randomly selected plants from each unit were marked and used to estimate yield components – plant height (cm), stem (mm) and length (cm) of the pod, number of seeds and pods per plant, number seeds per pod, 1000 seed weight (g) and seed weight per plant (g). The next statistical analyses were used to process the experimental data: genetic and phenotypic correlations were calculated by two-factor analysis of variance (ANOVA), regression and path analysis 10. All the experimental data were statistically processed using the computer software GENES 2009.7.0 for Windows XR 11.
RESULTS:
Correlation Analysis: The phenotypic correlation is conditioned by the relationship among individual characters and the influence of environmental factors. The genotypic correlation is a function of the pleiotropic action of the genes involved and their related inheritance. Linked genes have additive, dominant and epistatic actions. It is generally regarded that the additive genes are of greatest value in breeding 12. Inbreeding aspect, the presence of correlations is particularly important because of the possibility of using them as important criteria to conduct the more efficient and simultaneous phenotypic selection of two or more traits 13.
For spring pea varieties, coefficients of phenotypic dependence for most of the traits (with the exception of seeds per plant) were lower as compared to the coefficients of genotypic correlations Table 1.
Statistically significant and robust positive phenotypic correlations Table 2 were observed between plant height with seed per plant (r=0.588) and pod length (r=0.411). Highly positive were the dependencies between plant height with pods per plant (r=0.780); pods per plant with seeds per plant (r=0.863) and seed weight per plant (r=0.796); seeds per plant with seed weight per plant (r=0.733). Positive but weak were the relationships between pod stem with pod length (r=0.391) and seeds per plant (r=0.296); pod length with 1000 seed weight (r=0.214). There are established similar dependencies between seed yield and tested quantitative traits as well as in the phenotypic correlation.
TABLE 1: GENETIC CORRELATION COEFFICIENTS AMONG THE CHARACTERISTICS IN PEA VARIETIES
| Characteristics | Plant
height |
Pods
per plant |
Pod
stem |
Pod
length |
Seeds
per plant |
Seed weight per plant | 1000 seed
weight |
| Pods per plant | 0.7984 | ||||||
| Pod stem | 0.5151 | 0.5401 | |||||
| Pod length | 0.4479*** | 0.518 | 0.4334 | ||||
| Seeds per plant | 0.6124*** | 0.8662 | 0.3227 | 0.4959 | |||
| Seed weight per plant | 0.3892 | 0.7966 | 0.538 | 0.5973 | 0.7224 | ||
| 1000 seed weight | -0.3127 | -0.0027 | 0.3054*** | 0.2198 | -0.2352*** | 0.4561 | |
| Seed yield | 0.6715*** | 0.801 | 0.6924*** | 0.4872 | 0.4249 | 0.739 | 0.4419 |
High genetic correlations were detected between plant height with seeds per plant (r = 0.612) and pods per plant (r=0.798); pods per plant with seed per plant (r=0.866) and seed weight per plant (r=0.796); seed per plant with seed weight per plant (r=0.722). Moderate correlations were established between plant height with pod stem (r=0.515), pod length (r=0.447) and seed weight per plant (r=0.389); pod length with pod per plant (r=0.518) and seed weight per plant (r=0.597). Relatively low were correlations between 1000 seed weight with pod stem (r=0.305) and pod length (r=0.219); seeds per plant with pod stem (r=0.322).
Negative genetic correlation coefficients were detected between 1000 seed weight with plant height (r=-0.312) and seed per plant (r=-0.235). All tested features positively correlated with grain yield, although statistically proven are only with pod stem (r=0.692) and plant height (r=0.671).
Table 2: Phenotypic correlation coefficients among the characteristics in pea varieties
| Characteristics | Plant
height |
Pods
per plant |
Pod
stem |
Pod
length |
Seeds
per plant |
Seed weight
per plant |
1000
Seed weight |
| Pods per plant | 0.7805 | ||||||
| Pod stem | 0.5027 | 0.521 | |||||
| Pod length | 0.4112*** | 0.4943 | 0.3916 | ||||
| Seeds per plant | 0.5882*** | 0.863 | 0.2969 | 0.4799 | |||
| Seed weight per plant | 0.3849 | 0.7964 | 0.5028 | 0.5667 | 0.7334 | ||
| 1000 seed weight | -0.3048 | -0.005 | 0.3007*** | 0.2143 | -0.2255*** | 0.4401 | |
| Seed yield | 0.6518*** | 0.7875 | 0.6733*** | 0.477 | 0.4242 | 0.7178 | 0.4373 |
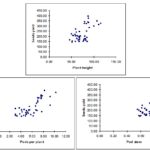
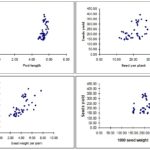
Path Coefficient Analysis: The path coefficient analysis Table 3, Fig. 1 revealed that pod stem (2991.134), seeds per plant (448.314) and pods per plant (99.261) were the productive components possessing the highest positive direct effects on seed yield.
They could be an important criterion for selecting desirable traits for genetic improvement of pea crop. The strongest indirect effects were by seeds per plant via plant height (14077.05), pod stem (12418.29) and 1000 seed weight as well as pod stem via pods per plant, pod length and seed weight per plant.
The highest total effect was observed in 1000 seed weight (197.31), plant height (115.60) and seeds per plant (19.00).
TABLE 3: PATH ANALYSIS FOR SEED YIELD OF PEA VARIETIES
| Traits | Indirect effect | Total effect | ||||||
| Plant
height |
Pods
per plant |
Pod
stem |
Pod
length |
Seeds
per plant |
Seed weight per plant | 1000 seed
weight |
||
| Plant height | -15.429 | 1012.464 | 2333.084 | -7122.900 | 14077.050 | -10290.800 | 1467.740 | 115.600 |
| Pods per plant | -1351.610 | 99.261 | 2572.375 | -7070.710 | 10759.530 | -7211.690 | 1495.777 | 7.600 |
| Pod stem | -1356.240 | 913.203 | 2991.134 | -7096.800 | 12418.290 | -8751.270 | 1420.374 | 0.810 |
| Pod length | -1228.180 | 774.237 | 2542.464 | -1304.560 | 10311.210 | -6509.430 | 1186.383 | 5.750 |
| Seeds per plant | -1339.270 | 754.385 | 2422.818 | -6940.260 | 448.314 | -5820.670 | 1258.389 | 19.000 |
| Seed weight per plant | -1283.720 | 714.681 | 2482.641 | -7005.490 | 9997.392 | -1350.500 | 1270.802 | 4.120 |
| 1000 seed weight | -1650.940 | 754.385 | 2333.084 | -7266.400 | 11118.180 | -6347.370 | 6.176 | 197.310 |
Fig. 1: Dependences between seed yield and quantitative traits
Regression Analysis: The results of carried out multiple regression analysis showed that the linear component in the regression of seed yield in respect to the investigated quantitative traits is significant and reliable Table 4. From the complex study of the traits is obtained model (1) that demonstrates the complex character of the change of productivity depending on the variation of investigated quantitative traits Table 5. The common type of the obtained equation of regression was:
(1) У = -294.32 + 0.824768* X1 + 39.37287* X2 + 50.62091* X3 + 19.71406* X4 - 4.70042* X5 -8.90222* X6 + 1.134677* X7
Where: Y – is seed yield; X1 – plant height; X2 – pods per plant; X3 – pod stem; X4 – pod length; X5 – seeds per plant; X6 – seed weight per plant; X7 – 1000 seed weight.
The applied analysis showed that for the formation of seed yield the highest influence had the pod stem (50.62) and pods per plant(39.37), followed by pod length (19.71). The graphical representation of the relations between the productiveness and studied quantitative components allows by adequate approaching to be obtained theoretical results and to be determined the basic regularity between these traits Fig. 2.
In the regression equation (1) the dependence between the seed yield and seed weight per plant (-8.90) was with the highest negative value. The grain yield increased by nearly 50.62 g if a unit increases the seed weight per plant. Lower was the influence of 1000 seed weight (1.13), plant height (0.82) and seeds per plant (-4.70) as at the last trait the coefficient of regression was negative.
TABLE 4: REGRESSION ANALYSIS (ANOVA) OF THE SEED YIELD REGARDING THE OTHER TRAITS
| df | SS | MS | F | Significance F | |
| Regression | 7 | 192516.8 | 27502.4 | 39.08469 | 3.31E-15 |
| Residual | 37 | 26035.49 | 703.6618 | ||
| Total | 44 | 218552.3 |
Table 5: Regression coefficients of the seed yield regarding the other traits
| Coefficients | Standard Errorr | t Stat | P-value | |
| Intercept | -294.32 | 106.4634 | -2.76451 | 0.008836 |
| Plant height | 0.824768 | 0.520535 | 1.584462 | 0.1216 |
| Pods per plant | 39.37287 | 7.894047 | 4.987666 | 1.47E-05 |
| Pod stem | 50.62091 | 53.10244 | 0.953269 | 0.34664 |
| Pod length | 19.71406 | 19.95159 | 0.988095 | 0.329524 |
| Seeds per plant | -4.70042 | 2.63861 | -1.7814 | 0.083058 |
| Seed weight per plant | -8.90222 | 10.78303 | -0.82558 | 0.414336 |
| 1000 seed weight | 1.134677 | 0.361075 | 3.142499 | 0.003292 |
FIG. 2: PATH ANALYSIS – EFFECT OF CO LINEARITY OF VARIABLES
Legend: 1-plant height; 2-pods per plant; 3-pod stem; 4-pod length; 5-seeds per plant; 6-seed weight per plant; 7-1000 seed weight
DISCUSSION: The presented data demonstrate that the seed yield in field pea is a complex trait and is the result of the combined effect of all seed yield components 14, 15. They also confirm the earlier reports that seed weight per plant is highly and positively correlated to a number of pods per plant and seeds per plant 16. Some researchers 17, 18 reported for a significantly positive relationship between seed weight and a number of seeds per pod, and other authors 18, 19 established that number of pods per plant exhibited a significant and positive association with seed weight per plant.
Goa and Ashamo 20 have also found a positive and highly significant dependence between seed yield and 1000 seed mass, seed weight per plant and pods per plant. The direct effect of any character on seed yield per plant gives an idea about effective selection that can be made to bring improvement. Therefore direct selection through these traits would be effective. Such similar results have been stated in cowpea 21. The indirect effect shows the interrelation of component characters towards contribution to yield. Thus, selection for a number of pods per plant, seeds per plant, and pod length would give good response in yield improvement 2. Number of seeds per plant showed a high direct effect on seed yield. Similar results are earlier reported by Ghobary 22.
In the breeding aspect, the high but non-significant correlations between tested traits can lead to increasing the values of one parameter and reducing the other one. The low and non-significant correlation is an indication of the presence of non-linearity interaction between traits 23.
CONCLUSION: In general, the genetic correlations for most of the traits in field pea have higher coefficients than the phenotypic correlations. The strongest positive phenotypic correlations were found between plant height with seed per plant (r=0.588), pod length (r=0.411) and pod per plant (r=0.780); pod per plant with seed per plant (r=0.863) and seed weight per plant (r=0.796); seed per plant with seed weight per plant (r=0.733). The highest genetic correlations were established between plant height with seed per plant(r = 0.612) and pods per plant (r=0.798); pods per plant with seed per plant (r=0.866) and seed weight per plant (r=0.796); seed per plant with seed weight per plant (r=0.722). The strongest and direct positive effects over grain yield had pod stem (2991.134), seeds per plant (448.314) and pod per plant (99.261). The highest positive indirect contribution was by seeds per plant via plant height (14077.05), pod stem (12418.29) and 1000 seed weight as well as by pod stem via pods per plant, pod length and seed weight per plant.
Based on the trait association and the path coefficients for seed yield and its components, it could be concluded that field pea breeders should pay attention to the traits as pod stem, seeds per plant, pods per plant and 1000 seed weight at selecting high-yielding genotypes.
ACKNOWLEDGEMENT: Nil
CONFLICT OF INTEREST: Nil
REFERENCES:
- Tyagi N, Singh AK, Rai VP, Kumar S and Srivastava CP: Genetic variability studies for lodging resistance and yield attributes in pea (Pisum sativum). Journal Food Legumes 2012; 25(3):179-182.
- Saxesena RR, Vidyakar V, Vishwakarma MK, Yadav PS, Meena ML and Lal GM: Genetic variability, correlation and path analysis studies in garden pea (Pisum sativum). The Bioscan (Supplement on Genetics and Plant Breeding) 2014; 895-898.
- Kumar NJ and Lavanya GR: Correlation coefficient analysis in field pea (Pisum sativum L). International Journal of Agricultural Science and Research 2014; 4(6): 211-214.
- Kumar B, Kumar A, Singh AK and Lavanya GR: Selection strategy for seed yield and maturity in field pea (Pisum sativum arvense). Global Journal of Crop, Soil Science and Plant Breeding 2013; 1(1): 129-133.
- Pratap A and Kumar J: Biology and breeding of food legumes. Kanpur: Indian Institute of Pulses Research, Edition 1st, 2011.
- Sood M and Kalia P: Gene action of yield-related traits in garden pea (Pisum sativum). Journal of Breeding and Genetics 2006; 38(1): 1-17.
- Togay N, Togay Y, Yildirim B and Dogan Y: Relationships between yield and some yield components in pea (Pisum sativums sparvense L.) genotypes by using correlation and path analysis. African Journal of Biotechnology 2008; 7(23): 4285-4287.
- Espósito MA, Martin E, Cravero V, Liberatti D, López A, Sebastián F and Cointry E: Relationships among agronomic traits and seed yield in pea. Journal of Basic & Applied Genetics 2009; 20(1): 1-8.
- Kalapchieva S: Model for breeding of high productive garden pea varieties. Plant Science 2013; 50(3): 73-76.
- Williams WA, Jones MB and Demment MW: A concise table for path analysis statistics. AgronomyJournal 1990; 82: 1022-1024.
- Cruz CD: Programa Genes: Biometria. Version 7.0. University of Federal Viçosa, Viçosa, Brazil, Edition 1st, 2006.
- Zhelyazkov E and Tsvetanova Y: Rakovod stvozaupraz-hneniyapo genetika. Trakian University Press, Stara Zagora 2002. [In Bulgarian].
- Juchenko AA, Nesterov V and Andryushenko VK: Mathematical modeling in the optimization of breeding-genetic researches. Kishinev 1980.
- Sardana S, Mahajan RK, Gautam NK and Ram B: Genetic variability in pea (Pisum sativum). SABRAO Journal of Breeding and Genetics 2007; 39: 31-41.
- Sultana ZAK, Islam MA, Hasan Mitu MK and Mian MAK: Genetic variability and character association in garden pea (Pisum sativum spp. hortense) genotypes. 2009;http://aminulgpb.faculty.bsmrau.edu.bd/files/2009/07/Zakia-paper _Genet-Var-BJAR.pdf
- Kosev V and Mikić А: Assessing relationships between seed yield components in field pea (Pisum sativum) cultivars by correlation and path analysis. Spanish Journal of Agricultural Research 2012; 10(4): 1075-1080.
- Usmani MG and Dubey RK: Association and path coefficient analysis in field pea (Pisum sativum averse). Advanced in Plant Science 2007; 20(2): 603-607.
- Singh A, Singh S and Babu JDP: Heritability, character association and path analysis studies in early segregating population of field pea (Pisum sativum arvense). International Journal of Plant Breeding and Genetics 2011; 5(1): 86-92.
- Jitendra K, Ashraf N and Pal K: Variability and character association in garden pea (Pisum sativum subsp. hortense Asch. And Graebn). Progressive Agriculturists 2010; 10(1): 124-131.
- Goa Y and Ashamo M: Evaluation of field pea (Pisum sativum) genotypes performance for yield and yield components at five growing environments of Southern Ethiopia. Current Research in Agricultural Sciences 2014; 1(3): 65-76.
- Manggoel W, Uguru MI, Ndam ON and Dasbak MA: Genetic variability, correlation and path coefficient analysis of some yield components of ten cowpeas [Vigna unguiculata (L.) Walp] accessions. Journal of Plant Breeding and Crop Science 2012; 4(5): 80-86.
- Ghobary HM: Study of Relationship between yield and some yield components in garden pea (Pisum sativum) by using correlation and path analysis. Agricultural Research 2010; 36: 351-360.
- Tsenov N, Kostov K, Gubatov T and Peeva V: Study on the genotype × environment interaction in winter wheat varieties. I. Grain quality. FieldCropsStudies2004; 1: 20-29.
How to cite this article:
Georgieva N, Nikolova I and Kosev V: Association study of yield and its components in pea (Pisum sativum L.). Int J Pharmacognosy 2015; 2(11): 536-42. doi link: http://dx.doi.org/10.13040/IJPSR.0975-8232.IJP.2(11).536-42.
This Journal licensed under a Creative Commons Attribution-Non-commercial-Share Alike 3.0 Unported License.
Article Information
3
536-542
651
2141
English
IJP
N. Georgieva *, I. Nikolova and V. Kosev
Department of Technology and Ecology of Forage Crops, Institute of Forage Crops, Pleven, Bulgaria.
imnatalia@abv.bg
07 September 2015
19 November 2015
28 November 2015
10.13040/IJPSR.0975-8232.IJP.2(11).536-42
30 November 2015