MOLECULAR BEACONS: FUNDAMENTAL ASPECTS AND APPLICATIONS
HTML Full TextMOLECULAR BEACONS: FUNDAMENTAL ASPECTS AND APPLICATIONS
Neelam Verma * 1, Navpreet Kaur 1 and Pawan Krishan 2
Department of Biotechnology 1, Department of Pharmaceutical Sciences and Drug Research 2, Punjabi University, Patiala - 147002, Punjab, India.
ABSTRACT: Molecular Beacons (MBs) are DNA hairpins that are widely used as fluorescent probes. The inherent signal transduction mechanism let them differentiate between target and non-target oligonucleotides that differ even on a single base. The stem-loop structure consists of donor and acceptor moieties conjugated to both ends thus lying nearby, which leads to FRET. On hybridization to target sequences that hairpin opens up, restoring fluorescence. MBs technology can be applied to a wide range of fields from genetic screening, biochip development, biosensor construction and even mRNA expression in living cells. Recent developments consist of modifications to improve not only the stability of MB in living cells but also lowering signal to noise ratio.
| Keywords: |
DNA hairpin, Fluorescent probes, Molecular genetics, SNP, DNA, RNA
INTRODUCTION: Single-stranded hairpin oligonucleotides probes named as molecular beacons were designed by Tyagi and Kramer in 1996 which indicate the presence of target nucleic acids present in homogenous solutions. Molecular beacons belong to a novel class of DNA probes that are widely used in the field of biology, biotechnology, chemistry and medical sciences for bio-molecular recognition 1. The uniqueness of these probes lies not in their functionality but also in ease of synthesis and their molecular specificity. One of the most potent advantages is their structural tolerance to various modifications 2. Molecular beacons are hairpin oligonucleotides with fluorophore and quencher coupled to opposite ends. Molecular beacons 3 indicate the presence of specific nucleic acids in homogenous solutions.
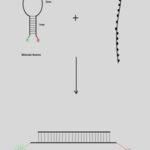
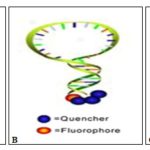
These molecular switches are in off mode (closed conformation) when the target molecule is absent. Here the complementary ends of the molecular beacon hybridize to give a classic hairpin conformation. This close structure of the probe provides not only low background fluorescence in the target unbound form but also provides high selectivity in comparison to linear probes. Formation of a hairpin loop structure brings fluorophore in proximity to quencher resulting in contact quenching 4.
FIG. 1: HYBRIDIZATION OF MOLECULAR BEACON TO TARGET SEQUENCE 3
Introduction of the pre-synthesized target molecule or PCR amplified DNA targets to the mixture results in the formation of beacon to target-hybrid (duplex). Formation of hybrid is termed as on mode as fluorophore and quencher are far apart resulting in no more quenching and thus fluorescence occurrence Fig. 1. The loop portion is the sensing element while stem along with both fluorophore and quencher are physiochemical transducer. Thus, either or both stem and loop portion of molecular beacons can be engineered to achieve the best possible performance of molecular beacon based biosensor.
2. Designing of Molecular Beacon: DNA base pairing is an operating principle of molecular beacons. The loop portion should consist of 15-20 nucleotides and fully complementary to the target molecule. The stem portion should be rich in G/C content and may contain 4-7 base pairs. The assay selectivity improves when longer stems are used while hybridization rates decline. To ensure efficient fluorescent quenching the melting temperature of the stem should be 5 °C higher than assay temperature (http.//www.molecular-beacon.com/MB_SC_design.html). For better allele differentiation in SNP specific assays, the interrogated position should be complementary to nucleotide close to the middle position of the loop sequence. To reduce the background noise of assay, quantum yield and quenching efficiency of fluorophore and quencher respectively should be taken into account 5.
The melting temperature can be adjusted (i) monitoring GC content of stem, (ii) varying the length of the stem, (iii) varying length & little presence of GC content in a loop. The melting temperature of probe-target hybrid can be predicted by GC rule, which is the basic principle of most online probe design programs such as Oligo 6.0 (Molecular Biology Insight, Inc., Cascade, Co, USA); Primer express, Primer Premier 6 prediction should be made before the addition of stem sequences. Stem formation is a consequence of a series of intramolecular hybridization event; thus its melting temperature can’t be predicted by percent GC rule. Instead, DNA folding programs such as Zuker folding program (http://www.bioinfo.rpi.edu/ applications/mfold/.) can be used 7.
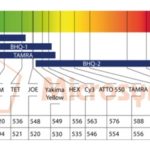
Formation of unwanted 2° structures, desolvation of the close proximity of fluorophore and quencher due to certain structures are indicated by such folding programs. Even nucleotides too exhibit a variable degree of quenching in order of G>A>C>T so their position just before the fluorophore should be wisely chosen. Different pairs of fluorophore and quencher can be used in Fig. 2.
FIG. 2: SCHEMATIC OF THE USE AND INTERPLAY OF COMMON FLUOROPHORES AND QUENCHERS 8
2.1. Fluorophore: Different fluorophore dyes have been tested to date. Among the range of dyes that justified their efficiency depending upon the quencher, group are EDNAS [5-(2’-aminoethyl) amino naphthalene-1-sulfonic acid, fluorescein [Fam], Tet [Tetrachloro-6-carboxyfluorescein], Hex [Hexachloro 6- carboxy fluorescein], TAMRA [Tetramethylrhodamine] and Rox [5-carboxyrhodamine-X].
2.2. Quencher: It captures the light energy from the fluorophore and dissipates it as heat, a phenomenon known as quenchers. The most commonly used quencher, i.e. Dabcyl[4-(4’-dimethyl aminophenylazo] benzoic acid. It is a neutral and hydrophobic molecule that serves as a universal quencher for a range of fluorophore 8. The light emitted by fluorescein is best quenched by dabcyl, but for others that emit longer wavelength, its quenching efficiency shortfalls to 93-98%.
The metal used as a quencher in molecular beacon analysis has bloomed a new era. Gold nanoparticles (1.4nm) 9 and use of gold chips 10 have been reported. Molecular beacons with gold as quencher offer great sensitivity to single nucleotide mismatch of DNA sequence.
3. Fundamental Aspects of Molecular Beacons:
3.1 Energy Transfer and Signal Transduction Mechanism: There are different energy transfer mechanisms for signal transduction in molecular beacons. Major categories are (i) Dynamic Quenching, (ii) Static fluorescence quenching. Dynamic quenching inculcates Forster transfer (RET or FRET) and Dextar transfer (collision quenching or ē transfer quenching). RET is a result of long-range dipole-dipole interactions between donor and acceptor and is without the release of a photon.
Here energy transfer rate depends upon the extent of spectral overlap between emission of the spectrum of donor and absorption spectrum of the acceptor. Förster distance is a distance at which RET occurs with 50% efficiency is typically in the range of 20-70Å. The mechanism to be followed depends upon the stem sequence and chromophores and linkers used. Ground state complexes are formed in static quenching. Also known as contact quenching, this was the original signal transduction mechanism for the molecular beacons.
The term contact quenching elaborates the proximity of a donor fluorophore to a non-fluorescent acceptor. Here most of the transferred energy is dissipated by a non-radiative process that is dissipated as heat 11, 12, 13.
3.2. Thermodynamics of Molecular Beacons: To state the thermodynamics of molecular beacons simple all or nothing [or “on”/ “off”] model has been used as an approximation of initial and final states of the reaction. Experimental data that supported this model showed the melting temperature of perfectly matched Target: MB. The model also predicted the high selectivity when the mutation was positioned at the center in loop region 5, 12, 14.
With the temperature change the phase of MB changes from duplex to hairpin to the random coil. Hybridization occurs in random coil stage. The selectivity of MB increases as stem length is increased as a result of the difference in phase transition between perfect complementary helix and mismatch duplex is increased 5, 15. Because of three phases thermodynamic behavior, the difference in transition temperatures between the perfectly matched helix and mismatched duplex increases contributing to enhanced selectivity of MBs. The selectivity of MB can be defined as the difference of phase transition temperature of perfectly matched Target: MB to phase transition temperature of mismatched Target: MB. There are 3 phases in which molecular beacon can exist: Hybridized to target, Hair-pin conformation and free as random coil 5, 14, 15, 16. It was proved that free energy of M.B is lower in stem-loop conformation that in random coiled state 8, 17, 18 (http://www.molecular-beacons. org/Introduction.html).
3.3. Kinetics of MBs: Probe and stem length doesn’t only affect the thermodynamic behavior of MBs but also the hybridization kinetics. Variation in either stem or loop alters the hybridization on-rate constants. Dual-labeled linear probes without the stem have 2x hybridization kinetic rate than MB with four stem bases 5. M B with longer probe length than shorter ones hybridizes more quickly to target sequences. Addition of single nucleotide to the stem of MB resulted in a 10-fold increase in on-rate constant.
Hairpin structure formation in an MB is driven by favorable free energy difference ΔGs. This ΔGs depends upon stem length Ls, stem sequence, ionic conditions and temperature 15, 19. Increase in stem length Ls increases ΔGs and thus results in the more stable stem-loop structure against thermal fluctuations. Also, the free energy difference ΔGp due to binding of the probe to its complementary target is monitored by probe length, probe sequence, and temperature. For a typical MB structure, ΔGp being larger than ΔGs, so stem-loop opens up on the molecular beacon to target hybridization. The conformational state of the molecular beacon at a given temperature depends upon ΔGp or ΔGs at a given ionic strength. Thus competition between ΔGp and ΔGs determines stability, specificity and hybridization kinetics.
3.4 Selectivity of MBs: MB probes can be immobilized onto surfaces and could be trapped at an interface, thus used for the development of biosensor 20, 21, 22, 23. Their excellent selectivity and high sensitivity make MB effective biosensor on gene chips and microarrays. The specificity of surface immobilized MB was elucidated by an experiment at a single molecule level by atomic force microscopy 24, 25. The rupture force required to separate MB and cDNA was detected and compared to that of linear DNA and its target. The results proved that molecular beacon was much more specific as their interaction force was 0.50 ± 0.11nN as compared to linear probe that was found to be 0.21 ± 0.08nN.
With an increase in the stem length of the MB discriminates target over a broader range of temperature is also increased. While the specificity of molecular beacon increases as probe length decreases but these alterations also leads to compromisation of hybridization rates 5.
3.5. Signal to Background Ratio of MBs: MB with longer stem has low background fluorescence than with short stem. The signal hike is observed when MB hybridizes to the target DNA. The success of phenomenon depends upon (i) fluorescence increase when the target and MB hybridize [S/N ratio] (ii) Dynamic range broadens, (iii) lower limit detection. To increase the S/N ratio either the fluorescent intensity of the fluorophore should be increased, or the background signal can be decreased.
4. Advantages of Molecular Beacons:
4.1. Sensitive Probe: It has a high signal to background ratio for real-time monitoring. Inherent fluorescent signal transduction mechanism in MB labels it as one of the major sensitive probes. 200 fold increase in the fluorescence intensity observed on its hybridization to target DNA under optimum conditions 3. MB can be used where it is not possible to isolate probe-target hybrids from the excess of unhybridized probes.
4.2. Real-Time Monitoring: This feature enables the synthesis of the nucleic acids to be monitored while the reaction is in progress 5, 26.
4.3. Sensitivity: M.B’s are extra-ordinarily target specific and select their counter partner as specific as without mismatch of Single Nucleotide (SN) 3.
4.4. It is a simple and promising tool for the dia-gnosis of genetic disease and gene-therapy study 27.
4.5. The temperature range at which M.B hybridizes to its complementary target is also wide than the mismatch ones 28.
5. Applications:
5.1. Real-Time Analysis: Real-time analysis is one of the most accurate and sensitive technique. MBs are one of the most attractive systems for a variety of bio-analytical applications in the world of biotechnology, biochemistry and biomedical sciences 29, 30, 31. Real-time analysis is one of the most attractive and sensitive means for quantitation of products. As the name illustrates it is monitoring of reaction products while the reaction is in progress. MB is used in collaboration of real-time assays because of better sensitivity and higher specificity than other assays 3 (http://www. molecular-beacons.org/Introduction.html).
DNA Detection: As the number of PCR cycles increases the high multiples of the target are produced which hybridizes with MBs at the annealing stage. This not only authenticates the formation of amplified product at high speed but also decrease the risk of contamination than other time-consuming processes such as gel electrophoresis and southern blotting. In 1998, Oritz et al., 32 developed surface probes to which PCR amplified amplicons were added. Verma and co-workers developed DNA based biosensor for hypertensive SNP (rs699) in 2016 and verified by applying it to the Punjabi population in 2017.
5.2. Real-Time Intracellular Imaging with Engineered Molecular Beacons: To yield important information about functional genomics numbers of probes have been developed to detect intracellular activities (such as synthesis, transport, and distribution) of mRNA in living cells 33. These inculcate FRET probes 34, GFP-fused RNA binding proteins 35, 36, 37, 38, Quenched Autoligation Probes 39, Nucleic acid stain SYTO14 40 and Molecular beacons 41, 42. The analyte can be detected without prior separation of unbound analytes.
Gene Expression Analysis: MB provide a unique and exquisite system for RNA analysis in living cells because of their higher selectivity (Single mismatch recognition), sensitivity (due to detection at a sub-nanomolar range, also low copy number of RNA) and real-time detection of mRNA as it has less lifetime. Various studies show that MBs don’t have deleterious effect on the living cells and can be used to monitor gene expression. Most important issues to consider are (i) designing of molecular beacon loop to the mRNA structure and (ii) Effective delivery of MBs into the system. The basic criteria for designing of MB are the prediction of the 2° structure of the mRNA. Moreover, the target region that is chosen should have a high probability of single-strandedness to promote efficient hybridization between MB and mRNA. For effective delivery various renowned such as electroporation 43 peptide assisted delivery 44, micro-injection 27, reversible permeabilization 27, streptomycin-O 45, standard bulk electroporation (Desai et al., 2013), microporation 46 and gold nanoprobes 47 can be chosen. The micro-injection delivers efficiently to a single cell but has low throughput. Electroporation and reversible permeabilization are high through-put techniques result in losses of cell integrity. In peptide assisted delivery peptide is conjugated with the probe which adds complexity and cost of the probe. Recently Nano Fountain (NFP) electroporation was reported. It delivers MBs into single cells without disturbing adhered cells. DNA based beacon detected glyceraldehyde 3- phosphate dehydrogenase and RNA based beacon detected green fluorescence protein mRNA. Also, time-dependent detection of mRNA expression is feasible by transfecting the same cell at a different time interval (throughout 24 h) 48.
Moreover, the incubation period is required for delivery processes which may affect the hybridization dynamics. After the effective entry of DNA into the cell, strategies to facilitate endosomal escape may be required for peptide conjugated probes because false positive signals may be generated. Earlier in-vivo detection was focused on target: RNA sequence hybridization rather than localization and distribution 49, 50. MB with oskar mRNA in oocytes of Drosophila melanogaster was investigated for its distribution and transport by Tyagi and co-workers 51 in 2003. Here two MBs were designed for adjacent regions of mRNA and FRET signal would occur only when both MBs were hybridized to mRNA. In this method, mRNA distribution was not only visualized, but its migration was also tracked. Bao and co-workers demonstrate the specific localization of both glyceraldehyde 3-phosphate dehydrogenase and K-ras in mitochondria using MB in conjunction with the fluorescent mitochondrial stain as the fluorescence of latter could be resolved spectrally.
The concentration of MB was resolved by attaching tRNA to the extended arm of MB than eliminating background fluorescence that was generated due to sequestration and opening up of MB in the nucleus. As a result, tRNA-coupled MB had longer residence time hence suitable for intracellular imaging 42. MB conjugated macromolecules and nanoparticles also eliminated the MB concentration in the nucleus 33, 52.
5.3. Neuron Cell Genomics: The extreme diversity, the complexity of the nervous system and mechanism underlying brain functions remain unsolved even after many years of extensive research 50. MB might prove as a sound answer to the mystery and thus help to gain insight into neural networks. Functions of single neurons can be investigated by real-time monitoring of the expression of living cells. mRNA of living neuron cells was studied with MB, but unfortunately, a high fluorescence was obtained soon after MBs were microinjected which might be the result of either protein binding or nuclease digestion of MBs 25. To increase the stability of MB in cells new form of MBs was synthesized known as Locked Nucleic Acids that have rigid structure 53, 54.
5.4. Triplex DNA Detection: Molecular Beacons have been used to study the kinetics of triplex DNA formation 59 and could be used to monitor DNA and RNA interaction. IN disease/disorder relevant genes promoter site is inhibited by the triplex DNA formation which could be analyzed in real-time to facilitate pharmacogenomics and anti-gene therapeutic strategies.
5.5. Molecular Engineering of MBs: To prevent non-specific protein binding and nuclease digestion, MB needs to be optimized before they can be used to monitor mRNA expression. This optimization not only enhances its specificity but also increases their residence time.
5.5.1. Increasing Sensitivity of Molecular Beacons: Conventional molecular beacons either report abundant or amplified gene products. Thus target sequences with low copy number have always been a big challenge. The sensitivity of molecular beacons can be enhanced by either increasing the fluorescence intensity of the fluorophore or improving the quenching efficiency of the quencher. Most of the strategies for improving the sensitivity of MB are focused on fluorescent intensity increment because the background signal from the cellular compartment and environment always affect the efficiency of the quencher. Super Quenching is achieved using a molecular assembly of multiple quenchers 55. Not only quenching efficiency of DABCYL for FAM hiked from 92.9 to 99.7% by using three DABCYL molecules instead of one but also signal to background ratio increased significantly from 14 to 320 Fig. 3.
FIG. 3: A) CHEMICAL STRUCTURE OF THREE DABCYL ATTACHED TO THE OLIGONUCLEOTIDES, B) MOLECULAR BEACON WITH ONE FLUOROPHORE AND SUPERQUENCHERS, C) COMPARISON OF QUENCHING IN SINGLE, DOUBLE AND TRIPLE ATTACHED QUENCHERS 55
5.5.2. Conjugated polymers (CPˉ) and their effect on sensitivity: CPˉ is spˉ or sp2ˉhybridized polyunsaturated macromolecules that exhibit photoluminescence with high quantum efficiency 56, 57. CPˉ are million-fold more sensitive to fluorescence compared to low molecular analogs thus demonstrating super quenching effect 25, 58. Using these fluorescent dyes such as Poly (phenylene ethylene) (PPE) has high fluorescence quantum yields when dissolved in the aqueous medium. Oligonucleotides were immobilized onto glass support that has pores of controlled size.
PPE dye molecules were directly polymerized onto oligonucleotides. The fluorescence of PPE was compared to other organic dyes such as Cy3, TMR, FAM, and Alex Fluor 488 and PPE was 20 times brighter than other dyes. Moreover, PPE neither interfered with the hairpin structure of molecular beacon nor with its hybridization rates 26.
5.5.3. Background Fluorescence Reduction: Incomplete quenching occurs due to the variety of reasons (i) Interruption of stem structures in traditional MBs, (ii) Non-specific binding of proteins or degradation by nucleases may generate false fluorescent signals (iii) imperfect quenching of fluorescence by quencher even when probe and quencher are nearby. And this occurs due to complicated cellular environment and thermodynamic conformation. To improve the signal to background ratio number of quencher can be increased. Yang et al. achieved high specificity and sensitivity by the assembly of a different number of quenchers on the stem of MBs 53. Introduction of multiple quenchers not only improves the absorption efficiency but also increases the probability of dipole-dipole coupling between the quencher and fluorophore.
5.5.4. Enhancing Biostability: Intracellular nuclease degradation and non-specific protein binding can be reduced by introducing artificial nucleotides. Artificial Nucleotides relies on artificially expanded genetic information (Aegis). Aegis [dZ:dP {6-amino-5-nitro-3-(1’beta-D-2’deoxyribofuranosyl)-2(1H)-pyridone: 2-amino-8-(1’-beta-D-2’-deoxyribofuranose)-imidaza(1,2-a)-1, 3,5-triazin-4(8H)-one}] pair was synthesized and incorporated into stem of molecular beacon which showed excellent resistance to enzymatic activity of nucleases in cell.
Also to enhance the stability of molecular beacon, Locked Nucleic Acids (LNA) were synthesized 53 by using methylene bridge connecting the 4’carbon and 2’ oxygen of the ribose. The LNA-LNA duplex has a stronger binding and stable at 95 °C. Also, LNA modified bases are better in single base mismatch detection. Moreover, they can also resist interference by non-specific DNA binding proteins and nucleases.
5.6. Molecule Interaction Study (Small Molecules and Protein): MBs are sensitive probes to monitor protein and enzymatic interactions with DNA 29, 60, 61. The molecular beacons can recognize proteins with E. coli single-stranded DNA binding proteins (SSB). SSB induces the conformational change in MB that affects the binding constant and binding stoichiometry which was measured by fluorescence enhancement.
Interaction of different nucleases such as S1 nuclease, DNase and mung bean nuclease with ssDNA was monitored with the help of MBs. The cleavage of MB by nuclease led to destabilization stem duplex and consequent fluorescence restoration 61. Various studies report the use of MB to study DNA ligation and phosphorylation 61, 62, 63, 64. Two short oligo sequences which have lower melting temperature than molecular beacon (as M.B remains intact) were designed complementary to the two adjacent zones on the loop of M.B. When introduced they hybridize to complementary loop sequence leading to nick in between oligo pairs. When DNA ligase was introduced it catalyzes the junction thus synthesizing the longer DNA sequence. Entire process leads to stem separation; hence fluorescence is restored 61, 62, 63. T4 DNA ligase and DNA ligase from E. coli have been explored with the help of M.B. Activity of DNA ligase are affected by metal ions, small bio-molecules and drugs 63.
NAD and ATP can also be investigated using MBs. Two short oligonucleotides complementary to the loop of Molecular Beacon were designed. Here DNA ligase repairs nick in the absence of the NAD. Assay thus reported was ten times faster 62, 65.
5.7. Molecular Beacons as Biosensors: The use of molecular beacons as biosensors is expanding rapidly 20, 21, 66, 67, 68, 69. MB should be immobilized onto a solid surface with high efficiency and must be optimized for use at the liquid-solid interface. For immobilization of MBs onto surface, either biotin-avidin interaction or thiol-gold linkage and amide bond could be employed. Different surfaces such as glass 20, gold 10, 22, polyacrylamide 53, agarose 53 have been used. All these surfaces not only provide efficient but also stable immobilization for a response, sensitivity, and reproducibility of MB based biosensors and biochips.
Background fluorescence generation in case of the biosensor is a big hurdle. Inculcation of space between the MB and the surface is one possibility 70, 71, 72. Poly T linker was also introduced as spacer though it improved little sensitivity. M.B attached to the surface of functionalized hydrophilic gel film of agarose, polyacrylamide also showed lower background fluorescence, higher sensitivity, faster response and better selectivity 21, 53. However, MB immobilization onto gold surface not only lowered the background fluorescence but also enhanced hybridization with target 10, 22.
Apart from fluorescence analysis enzymatic signal detection and electrochemical electron transfer have been developed. The reporter is an enzyme or electrochemical agent linked to the free end of MB. When MB hybridizes to target nucleic acid signal is released by a reporter (Conformation of reporter changes) 73, 74, 75.
Beads functionalized with MB have been used for multiplexed analyte detection. Here MB’s are immobilized onto microspheres that were entrapped within array of wells that was etched upon optical fiber 75. This array had a fast response time led to accurate analysis of multiple genetic mutations. Flow cytometry is another high throughput technique that could be combined with MB coated microsphere arrays to detect nucleic acids. Based on size and color coding signals from these were differentiated. Such assays that are simple, fast and accurate help in genetic analysis and genotyping for disease diagnosis and therapy.
Specific RNA or DNA targets in living cells can be detected by ultrasmall optical-fiber probes that provide not only extreme sensitivity but also enable spatial resolution.
5.8. Biomedical and Clinical Application: MBs have a range of clinical applications due to their outstanding properties. MBs combined with RT-PCR can detect gene mutations, pathogens, and other human diseases.
TABLE 1: MOLECULAR BEACON-BASED ASSAYS FOR HUMAN, FOOD AND ENVIRONMENTAL PATHOGEN (hMPV: HUMAN METAPNEUMOVIRUS, HPIV: HUMAN PARAINFLUENZA VIRUS, RSV: RESPIRATORY SYNCYTIAL VIRUS) 76
| Organism | Amplification Scheme |
| HIV | NASBA, PCR |
| HTLV | PCR |
| Oncogene HPV | NASBA, PCR |
| Mycobacterium tuberculosis | PCR |
| hMPV | NASBA |
| RSV | NASBA, PCR |
| Enterovirus | NASBA |
| Influenza virus | NASBA,PCR |
| Entamoeba histolytica | PCR |
| Methicillin-resistant Staphylococcus | PCR |
| Hepatitis B | NASBA, PCR |
| HPIV | NASBA |
| West Nile Virus | NASBA, PCR |
| Candida dubliniensis | PCR |
| Scedosporium | PCR |
| Pan-bacteria | NASBA, PCR, melting analysis |
| Pan-fungi | NASBA, PCR |
| Mycobacterial species | PCR, melting analysis |
| Pneumonia-causing agents | PCR |
| Chlamydophila pneumonia | NASBA |
| Multiple bacteria in bloodstream | PCR |
| Lyme disease spirochetes | PCR |
| Plasmodium | NASBA, PCR |
| Salmonella | PCR |
| Escherichia coli | NASBA |
| Listeria | NASBA |
| Environmental pathogens | |
| Salmonella | PCR |
| Escherichia coli | PCR |
| Baylisascaris procyonis | PCR |
| Vibrio cholerae | NASBA, PCR |
| Bacillus anthracis | PCR |
Use of Molecular beacons for cancer diagnostics not only provides reliable insight into carcinogenesis but also is promising method gene-expression detection in malignant tissues. Breast cancer gene BRCA-1 was detected in solution with LOD of 70nM 77. Medley along with his co-workers reported the expression of multiple genes inside a single breast carcinoma cell using MBs labeled with different-colored fluorophores. Survivin is an inhibitor of an apoptosis protein family and is a promising tumor marker as it is overexpressed in tumor cells. Tumor cells were stimulated with epidermal growth factor, treated with docetaxel led to overexpression of survivin gene which was real-time detected with MBs 78.
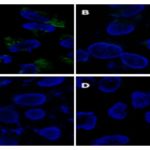
The progression of various cancers such as lung and ovarian cancers has been associated with Exon 2 deletion in aminoacyl tRNA synthetase complex-interacting multifunctional protein 2 (AIMP2). MBs with RT-PCR allowed the sensitive detection of the AIMP2-DX2 mutation. Dual-conjugated liposomes with molecular beacon and folate enabled fluorescence imaging of cancer cells harboring the AIMP2-DX2 mutation with high resolution 79. Cancer-associated fibroblasts of human epithelial carcinomas express a cell surface serine protease, i.e. Fibroblast activation protein (FAP) which is usually not expressed in normal fibroblasts, normal tissues, and cancer cells.
FIG. 4: FLUORESCENCE IMAGING OF THE HeLa CELLS USING LIPOSOMES CONJUGATED TO MOLECULAR BEACON (A) AIMP2-DX2 MUTANT FLUORESCED WITH DUAL-CONJUGATED LIPOSOMES (FOLATE AND MOLECULAR BEACONS). (B) LIPOSOMES CONJUGATED ONLY WITH MOLECULAR BEACONS. (C) LIPOSOMES CONJUGATED WITH RANDOM MOLECULAR BEACONS AND FOLATE. (D) LIPOSOMES CONJUGATED WITH RANDOM MOLECULAR BEACONS 79
Here peptide sequence (TSGPNQEQK) specific for FAP acted linker between black hole quencher and molecular beacon (FAP-PPB). When introduced to FAP in both in-vitro and in-vivo it cleaved the linker peptide restoring fluorescence. Thus, it efficiently detected epithelial cancer 80. Biosensor based on label-free molecular beacon was designed by Cao Q and co-workers in 2015 that studied the expression of Human Immuno Deficiency Virus (HIV), Hepatitis B virus (HBV) and Human T-Lymphotropic Virus Type I (HTLV).
CONCLUSION: Introduction of MBs to the molecular world has revolutionized real-time studies. They not only consist of highly transduction mechanism but are also flexible to chemical modifications. Since, these little miracles could detect both DNA and RNA, this has led to their broad spectrum application both cell genomics and expression.
According to WHO, 35 million people worldwide were infected with HIV in 2013. It is the world’s infectious killer, and till 2013, 39 million people have died. Among various nucleic acid assays that have been developed for real-time expression studies of viruses, molecular beacons not only represent an alternative screening method which is rapid, specific and relatively inexpensive. A micro-injection of multiple MBs with different fluoro-phores into HIV infected human cells makes it possible to monitor the expression of multiple genes simultaneously. This potent application not only will provide wealth information about HIV infected cells but also in preventing this deadly disease. Diseases of civilization such as hypertension and diabetes became more apparent in the modern world with industrialization. These diseases are not only affected by the environment but also by genes. The risk of developing a particular disease may increase or decrease with genetic variations with which response to particular drug also changes. MBs may not only prove quick and specific biosensors for in hand detection of such diseases but also can be a breakthrough technology in the screening of available drug against a particular disease. Thus, MBs will uplift pharmacogenomics by providing rational means to optimize drug therapy with maximum efficacy of drug and minimum adverse effects.
ACKNOWLEDGEMENT: This work is supported by Department of Biotechnology [DBT], India, under the project entitled as “DBT-Punjabi University, Patiala, Interdisciplinary Life Sciences Programme for Advanced Research and Education.”
CONFLICT OF INTEREST: Nil
REFERENCES:
- Tong W, Wang K, and Drake T: Molecular beacons. Current Opinion in Chemical Biology 2004; 8: 547-53.
- Drake TJ and Tan W: Molecular beacon DNA probes and their bioanalytical applications. Appl Spectrosc 2004; 58(9): 269A-280A.
- Tyagi S and Kramer FR: Molecular beacons: probes that fluoresce upon hybridization. Nat Biotechnol 1996; 14(3): 303-08.
- Kolpaschchikov DM: An elegant biosensor molecular beacon probe, challenges and recent solutions. Scientifica 2012; Article ID 928783: 1-17.
- Tsourkas A, Behlke MA, Rose SD and Bao G: Hybridization kinetics and thermodynamics of molecular beacons, Nucleic Acids Res 2003; 31(4): 1319-30.
- Marras SA, Tyagi S and Kramer FR: Real-time assays with molecular beacons and other fluorescent nucleic acid hybridization probes. Clin Chim Acta 2006; 363(1-2): 48-60.
- Markham NR and Zuker M: DINA Melt web server for nucleic acid melting prediction. Nucleic Acids Res 2005; 33: W577-81.
- Tyagi S, Bratu DP and Kramer FR: Multicolor molecular beacons for allele discrimination. Nat Biotechnol 1998; 16: 49-53.
- Dubertret B, Calame M and Libchaber AJ: Single-mismatch detection using gold-quenched fluorescent oligonucleotides. Nat Biotechnol 2001; 19(4): 365-70.
- Du H, Strohsahl CM, Camera J, Miller BL and Krauss TD: Sensitivity and specificity of metal surface-Immobilized “molecular beacons.” Biosensor J Am Chem Soc 2005; 127(21): 7932-40.
- Lakoweiz JR: Principles of Fluorescence Spectroscopy, third ed., Springer US 1999.
- Tsourkas A, Behlke MA and Bao G: Structure-function relationships of shared-stem and conventional molecular beacons. Nucleic Acids Res 2002; 30(19): 4208-15.
- Wang Y, Li J, Jin J, Wang H, Tang H, Yang R and Wang K: Strategy for molecular beacon binding readout separating molecular recognition element and signal reporter. Anal Chem 2009; 81(23): 9703-09.
- Bratu DP, Cha BJ, Mhlanga MM, Kramer FR and Tyagi S: Visualizing the distribution and transport of mRNAs in living cells. Proc Natl Acad Sci USA 2003; 100(23): 13308-13.
- Bonnet G, Tyagi S, Libchaber A and Kramer FR: Thermodynamic basis of the enhanced specificity of structured DNA probes, Proc Natl Acad Sci 1999; 96(11): 6171-76.
- Tan L, Li Y, Drake TJ, Moroz L, Wang K, Li J, Munteanu A, Chaoyong JY, Martinez K and Tan W: Molecular beacons for bioanalytical applications. Analyst 2005; 130(7): 1002-05.
- Tyagi S, Marras SA and Kramer FR: Wavelength-shifting molecular beacons. Nat Biotechnol 2000; 18(11): 1191-96.
- Bratu DP, Catrina IE and Marras SA: Tiny molecular beacons for in-vivo mRNA detection. Methods Mol Biol 2011; 714: 141-57.
- Wallace MI, Ying L, Balasubramanian S and Klenerman D: Non-Arrhenius kinetics for the loop closure of a DNA hairpin, Proc Natl Acad Sci USA 2001; 98(10): 5584-89.
- Fang XH, Liu XJ, Schuster S and Tan WH: Designing a novel molecular beacon for surface-immobilized DNA hybridization studies. J Am Chem Soc 1999; 121: 2921-21.
- Wang H, Li J, Liu H, Liu Q, Mei Q, Wang Y, Zhu J, He N and Lu Z: Label-free hybridization detection of a single nucleotide mismatch by immobilization of molecular beacons on an agarose film. Nucleic Acids Res 2002; 30(12): doi: 10.1093/nar/gnf061, PMID: 12060699.
- Du H, Disney MD, Miller BL and Krauss TD: Hybridization-based unquenching of DNA hairpins on au surfaces: prototypical "molecular beacon" biosensors. J Am Chem Soc 2003; 125(14): 4012-13.
- Cederquist KB, Golightly RS and Keating CD: Molecular beacon-metal nanowire interface: effect of probe sequence and surface coverage on sensor performance. Langmuir 2008; 24(16): 9162-71.
- Jin Y, Wang K, Tan W, Wu P, Wang Q, Huang H, Huang S, Tang Z and Guo Q: Monitoring molecular beacon/DNA interactions using atomic force microscopy. Anal Chem 2004; 76(19): 5721-25.
- Wang K, Tang Z, James-Yang CJ, Kim Y, Fang X, Li W, Wu Y, Medley CD, Cao Z, Li J, Colon P, Lin H and Tan W: Molecular Engineering of DNA: Molecular Beacons. Angew Chem Int Ed Engl. doi: 1002/anie.200800370.
- Kushon S, Bradford K, Marin V, Suhrada C, Armitage BA, McBranch D and David W: Detection of Single Nucleotide Mismatches via Fluorescent Polymer Superquenching. Langmuir 2003; 19: 6456-64.
- Medley CD, Drake TJ, Tomasini JM, Rogers RJ and Tan W: Simultaneous monitoring of the expression of multiple genes inside of single breast carcinoma cells. Anal Chem 2005; 77(15): 4713-18.
- Tsourkas A, Behlke MA, Rose SD and Bao G: Hybridization kinetics and thermodynamics of molecular beacons. Nucleic Acids Res 2003; 31(4): 1319-30.
- Tan W, Wang K and Drake TJ: Molecular beacons. Curr Opin Chem Biol 2004; 8(5): 547-53.
- Goel G, Kumar A, Puniya AK, Chen W and Singh K: Molecular beacon: a multitask probe. J Appl Microbiol 2005; 99(3): 435-42.
- Marras SA: Selection of fluorophore and quencher pairs for fluorescent nucleic acid hybridization probes. Methods Mol Biol 2006; 335: 3-16.
- Ortiz E, Estrada G and Lizardi PM: PNA molecular beacons for rapid detection of PCR amplicons. Mol Cell Probes 1998; 12(4): 219-26.
- Chen AK, Rhee WJ and Bao G: Delivery of molecular beacons for live-cell imaging and analysis of RNA. Methods Mol Biol 2011; 714: 159-74.
- Chen AK, Behlke MA and Tsourkas A: Avoiding false-positive signals with nuclease-vulnerable molecular beacons in single living cells. Nucleic Acids Res 2007; 35(16): PMID: 17702767.
- Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH and Long RM: Localization of ASH1 mRNA particles in living yeast. Mol Cell 1998; 2(4): 437-45.
- McDonald D, Vodicka MA, Lucero G, Svitkina TM, Borisy GG, Emerman M and Hope TJ: Visualization of the intracellular behavior of HIV in living cells. J Cell Biol 2002; 159(3): 441-52.
- Forrest KM and Gavis ER: Live imaging of endogenous RNA reveals a diffusion and entrapment mechanism for nanos mRNA localization in Drosophila, Curr Biol 2003; 13(14): 1159-68.
- Fusco D, Accornero N, Lavoie B, Shenoy SM, Blanchard JM, Singer RH and Bertrand E: Single mRNA molecules demonstrate probabilistic movement in living mammalian cells. Curr Biol 2003; 13(2): 161-67.
- Sando S and Kool ET: Imaging of RNA in bacteria with self-ligating quenched probes. J Am Chem Soc 2002; 124(33): 9686-87.
- Knowles RB, Sabry JH, Martone ME, Deerinck TJ, Ellisman MH, Bassell GJ and Kosik KS: Translocation of RNA granules in living neurons. J Neurosci 1996; 16(24): 7812-20.
- Bratu DP: Molecular beacons light the way: Imaging native mRNAs in living cells. Discov Med 2003; 3(19): 44-47.
- Mhlanga MM, Vargas DY, Fung CW, Kramer FR and Tyagi S: tRNA-linked molecular beacons for imaging mRNAs in the cytoplasm of living cells, Nucleic Acids Res 2005; 33(6): 1902-12.
- Golzio M, Rols MP and Teissié J: In-vitro and in-vivo electric field-mediated permeabilization, gene transfer, and expression, Methods 2004; 33(2): 126-35.
- Nitin N, Santangelo PJ, Kim G, Nie S and Bao G: Peptide-linked molecular beacons for efficient delivery and rapid mRNA detection in living cells. Nucleic Acids Res 2004; 32(6): PMID: 15084673.
- Nitin N, Rhee WJ and Bao G: Translation inhibition reveals interaction of 2'-deoxy and 2'-O-methyl molecular beacons with mRNA targets in living cells. Nucleic Acids Res 2009; 37(15): 4977-86.
- Desai HV, Voruganti IS, Jayasuriya C, Chen Q and Darling EM: Live-cell, temporal gene expression analysis of osteogenic differentiation in adipose-derived stem cell. Tissue Eng Part A 2013; 19(1-2): 40-48.
- Zhou Z, Zhang C, Qian Q, Ma J, Huang P, Zhang X, Pan L, Gao G, Fu H, Fu S, Song H, Zhi X, Ni J and Cui D: Folic acid-conjugated silica capped gold nanoclusters for targeted fluorescence/X-ray computed tomography imaging. J Nanobiotechnology 2013; 11: PMID: 23718865.
- Kang WM, Yavari F, Jaolandan MM, Vela GJP, Safi A, McNaughton RL, Parpoil V and Espinosa HD: Nanofountain probe electroporation (NFP-E) of single cells. Nano Lett 2013; 13(6): 2448-57.
- Perlette J and Tan W: Real-time monitoring of intracellular mRNA hybridization inside single living cells. Anal Chem 2001; 73(22): 5544-50.
- Drake TJ, Medley CD, Sen A, Rogers RJ and Tan W: Stochasticity of manganese superoxide dismutase mRNA expression in breast carcinoma cells by molecular beacon imaging. Chembiochem 2005; 6(11): 2041-2047.
- Bratu DP, Cha BJ, Mhlanga MM, Kramer FR and Tyagi S: Visualizing the distribution and transport of mRNAs in living cells. Proc Natl Acad Sci U S A 2003; 100(23): 13308-13.
- Tyagi S and Alsmadi O: Imaging native beta-actin mRNA in motile fibroblasts. Biophys J 2004; 87(6): 4153-4162.
- Wang L, Yang CJ, Medley CD, Benner SA and Tan W: Locked nucleic acid molecular beacons. J Am Chem Soc 2005; 127(45): 15664-65.
- Yang CJ, Medley CD and Tan W: Monitoring nucleic acids using molecular beacons. Curr Pharm Biotechnol 2005; 6(6): 445-52.
- Yang CJ, Lin H and Tan W: Molecular assembly of superquenchers in signaling molecular interactions, J Am Chem Soc 2005; 127(37): 12772-73.
- Hide F, Schwartz MA, Garchia D and Heeger AJ: Conjugated polymers are solid state laser materials. Synthetic Metals 1997; 91(1-3): 35-40.
- Wu CS, Peng L, You M, Han D, Chen T, Williams KR, Yang CJ and Tan W: Engineering molecular beacons for intracellular imaging. Int J Mol Imaging 2012; 501579. doi: 10.1155/2012/501579.
- Yang CJ, Lin H and Tan W: Molecular assembly of superquenchers in signaling molecular interactions. J Am Chem Soc 2005; 127(37): 12772-73.
- Antony T, Thomas T, Sigal LH, Shirahata A and Thomas TJ: A molecular beacon strategy for the thermodynamic characterization of triplex DNA: triplex formation at the promoter region of cyclin D1. Biochemistry 2001; 40(31): 9387-95.
- Fang X, Li JJ, Perlette J, Tan W and Wang K: Molecular beacons: novel fluorescent probes. Anal Chem 2000; 72(23): 747A-753A.
- Li JJ, Geyer R and Tan W: Using molecular beacons as a sensitive fluorescence assay for enzymatic cleavage of single-stranded DNA. Nucleic Acids Res 2000; 28(11): e52.
- Tang ZW, Wang KM, Tan WH, Li J, Liu LF, Guo QP, Meng XX, Ma CB and Huang SS: Real-time monitoring of nucleic acid ligation in homogenous solutions using molecular beacons. Nucleic Acids Research 2003; 31: e148.
- Liu LF, Tang ZW, Wang KM, Tan WH, Li J, Guo QP, Meng XX and Ma CB: Using molecular beacon to monitor activity of coli DNA ligase. Analyst 2005; 130: 350-57.
- Tang ZW, Wang K, Tan WH, Ma CB, Li J, Liu LF, Guo QP and Meng XX: Real-time investigation of nucleic acids phosphorylation process using molecular beacons. Nucleic Acids Research 2005; 33: e97.
- Ma C, Yang X, Wang K, Tang Z, Li W, Tan W and Lv X: A novel kinase-based ATP assay using molecular beacon, Anal Biochem 2008; 372(1): 131-33.
- Drake T and Tan W: Molecular Beacon DNA probes and their bioanalytical applications-, applied spectroscopy. Appl Spectrosc 2004; 58(9): 269A-280A.
- Chen AK, Rhee WJ and Bao G: Delivery of molecular beacons for live-cell imaging and analysis of RNA. Methods Mol Biol 2011; 714: 159-74.
- Epstein JR, Leung AP, Lee KH and Walt DR: High-density, microsphere-based fiber-optic DNA microarrays. Biosens Bioelectron 2003; PMID 12706560: 541-46.
- Stobiecka M and Chalupa A: Biosensors based on molecular beacons. Chemical Papers 2015; 69(1): 62-76.
- Li J, Tan W, Wang K, Xiao D, Yang X, He X and Tang Z: Ultrasensitive optical DNA biosensor based on surface immobilization of molecular beacon by a bridge structure. Anal Sci 2001; 10: 1149-53.
- Horejsh D, Martini F, Poccia F, Ippolito G, Di-Caro A and Capobianchi MR: A molecular beacon, a bead-based assay for the detection of nucleic acids by flow cytometry. Nucleic Acids Res 2005; 33(2): e13.
- Fan C, Plaxco KW and Heeger AJ: Electrochemical interrogation of conformational changes as a relentless method for the sequence-specific detection of DNA. Proc Natl Acad Sci USA 2003; 100(16): 9134-7.
- Bockisch B, Grunwald T, Spillner E and Bredehorst R: Immobilized stem-loop structured probes as conformational switches for enzymatic detection of microbial 16S rRNA. Nucleic Acids Res 2005; 33(11): e101.
- Graf N and Krämer R: Enzymatic amplification in a bioinspired, autonomous signal cascade. Chem Commun (Camb) 2006; 42: 4375-76.
- Steemers FJ, Ferguson JA and Walt DR: Screening unlabeled DNA targets with randomly ordered fiber-optic gene arrays. Nat Biotechnol 2000; 18(1): 91-94.
- Tyagi S and Kramer FR: Molecular beacons in diagnostics. F1000 Med Rep 2012; 4: 10.
- Culha M, Stokes DL, Griffin GD and Vo-Dinh T: Application of a miniature biochip using the molecular beacon probe in breast cancer gene BRCA1 detection. Biosensors and Bioelectronics 2004; 19: 1007-12.
- Peng XH, Cao ZH, Xia JT, Carlson GW, Lewis MM, Wood WC and Yang L: Real-time detection of gene expression in cancer cells using molecular beacon imaging: New strategies for cancer research. Cancer Research 2005; 65: 1909-1917.
- Jo SM, Kim YW, Jeong YS, Oh YH, Park KC and Kim HS: Rapid detection of exon 2-deleted AIMP2 mutation as a potential biomarker for lung cancer by molecular beacons. Biosensors and Bioelectronics 2013; 46: 142-49.
- Lo PC, Chen J, Stefflova K, Warren MS, Navab R, Bandarchi B, Mullins S, Tsao M, Cheng JD and Zheng G: Photodynamic molecular beacon triggered by fibroblast activation protein on cancer-associated fibroblasts for diagnosis and treatment of epithelial cancers, Journal of Medicinal Chemistry 2009; 52: 358-68.
- Guidelines for the use of fluorophores and quenchers, Mycrosynth.
- http://www.bioinfo.rpi.edu/applications/mfold/.
How to cite this article:
Verma N, Kaur N and Krishan P: Molecular beacons: fundamental aspects and applications. Int J Pharmacognosy 2019; 6(4): 117-27. doi link: http://dx.doi.org/10.13040/IJPSR.0975-8232.IJP.6(4).117-27.
This Journal licensed under a Creative Commons Attribution-Non-commercial-Share Alike 3.0 Unported License.